DOIng it Right! (DOIs for This Blog)
1This blog is an experiment. We want to share bits and pieces of our work; the reports we write, the presentations we hold, or the little discoveries we make, or even some first, water-testing investigations; and all the rest. It’s a documentation of what we do. Little bits of science, collected in the open, and sometimes even not that little.
 In the spirit of Open Science, wouldn’t it be great to acknowledge these little bits of science blog posts and have the option to refer to them in a scientifically sound way? Like… with DOIs, Digital Object Identifiers; the gold standard for referring to scientific work? Well, 🥁, the posts in this blog have DOIs now, including their metadata stored in a metadata repository!
In the spirit of Open Science, wouldn’t it be great to acknowledge these little bits of science blog posts and have the option to refer to them in a scientifically sound way? Like… with DOIs, Digital Object Identifiers; the gold standard for referring to scientific work? Well, 🥁, the posts in this blog have DOIs now, including their metadata stored in a metadata repository!
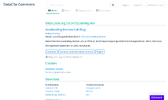 Thanks to help from our Forschunsgzentrum Library, we are able to use DataCite as DOI provider. I built a little Python tool which uses the DataCite API to register metadata of a blog post and mint a DOI. The DOI is shown in the header of each post, next to the license of the post (which is also new); the first part of the suffix of the DOI always containing
Thanks to help from our Forschunsgzentrum Library, we are able to use DataCite as DOI provider. I built a little Python tool which uses the DataCite API to register metadata of a blog post and mint a DOI. The DOI is shown in the header of each post, next to the license of the post (which is also new); the first part of the suffix of the DOI always containing xdvblg. I have released the Python tool as Open Source software as well, with a Zenodo DOI attached. It should be suitable for any other Jekyll-based blog as well!
I created DOIs retroactively for all previous blog posts, allowing us to link and refer to them a little bit more properly in scientific contexts2 from now on and have the metadata discoverable. Let’s see, if it sticks!
Read on for some technical details and design decisions.
DataCite is a service to store metadata of publications and create an optional DOI for it. Metadata can be viewed through their website or queried via APIs.
The Python tool, which I call doi-jekyll, is hosted on GitHub and released with an MIT license. Snapshots at Zenodo and according DOIs are automatically created for every GitHub release via the Zenodo GitHub integration. With this blog post I released v1.0!
doi-jekyll is a command line application which can be installed via pip3. It parses metadata of different locations within a Jekyll blog tree structure, assembles them as a validating instance according to the DataCite Metadata Scheme, submits the metadata to DataCite, and registers an auto-generated DOI. To build the metadata, data from an individual blog post (like title, license, abstract), from an author file (like name, ORCID ID), and from the blog itself (like blog title, blog DOI, but also API endpoint) are collected. The blog DOI4 is given as a Collection metadata of which every blog post inherits, creating a relationship between blog posts and the blog itself. The API with the latest Schema version is only available via an XML API (the JSON API is stuck on an older version, which doesn’t support the cool relational info). Because of this, the metadata is assembled in doi-jekyll as a Python dictionary and then internally converted to XML via xmltodict; a little bit of an extra effort, but working with Python dictionaries is so much easier compared to XML5.
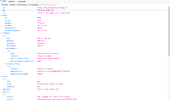 It took me a little bit of trial and error to assemble a validating metadata package which conforms to the DataCite Metadata Schema; luckily, DataCite has a test instance (called Fabrica Test) to fiddle around. While the interface of
It took me a little bit of trial and error to assemble a validating metadata package which conforms to the DataCite Metadata Schema; luckily, DataCite has a test instance (called Fabrica Test) to fiddle around. While the interface of doi-jekyll is made for this blog, it should work for any Jekyll blog; it has plenty of command line (and other) options to configure usage. A few examples: For testing, --dry-run allows to skip communication with DataCite, but do all the rest; and --skip-url will just register metadata at DataCite, but not mint a DOI. With --additional-metadata, further metadata can be specified to integrate into the to-be-uploaded metadata; an according key doi-additional-metadata in the YAML front matter of the post is available. To document and show some examples, the GitHub repository features an example Jekyll blog which has an example of all necessary files.
Let’s see where this weird journey of sciencifying blog posts leads us. The first person to place a xdvblg DOI reference in a paper gets a cupcake!
-
Alternative title of the post: DOIs “R” Us. It did not make the cut. ↩
-
Or as fancy shortlinks with attached metadata! ↩
-
Not from the Python Package Index, yet; one needs to use the GitHub URL directly. ↩
-
The parent DOI of the blog itself is https://doi.org/10.34732/xdvblg-mn,
mnlikemain, you know? ↩ -
Sigh, XML, amirite? ↩